These past few weeks I have continued running experiments comparing the calculated kmer frequency score and the folding energy of the sequence as it is mutated. Here the kmer frequency score has been maximized. Between experiments I have also been playing with my mutation function to try and make it more efficient. When running the experiments the higher the kmer frequency score got the more trouble the program had finding a synonymously mutated sequence that had an even higher score.
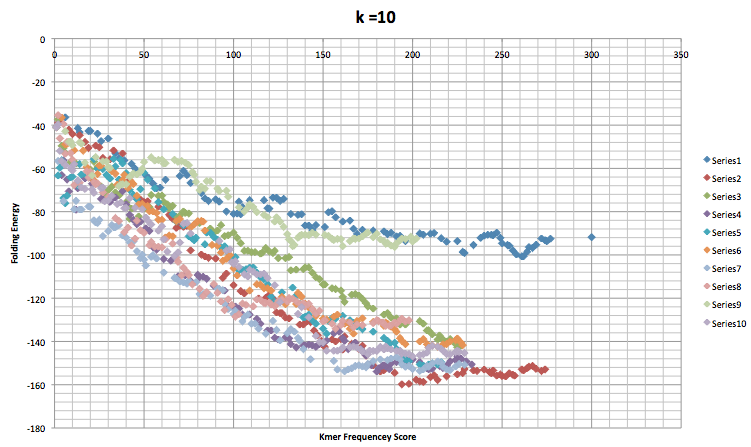
The graph below is the compiled data for when the kmer size was set to 10. Each different color refers to a different time the program was run. The legend on the right shows what color refers to what trial. Series 1 was the first time the program was run, series 2 the second time, and so on. Each run went through 100 accepted mutations and there are 10 runs shown below.
The graph below is the compiled data for when the kmer size was set to 10. Each different color refers to a different time the program was run. The legend on the right shows what color refers to what trial. Series 1 was the first time the program was run, series 2 the second time, and so on. Each run went through 100 accepted mutations and there are 10 runs shown below.
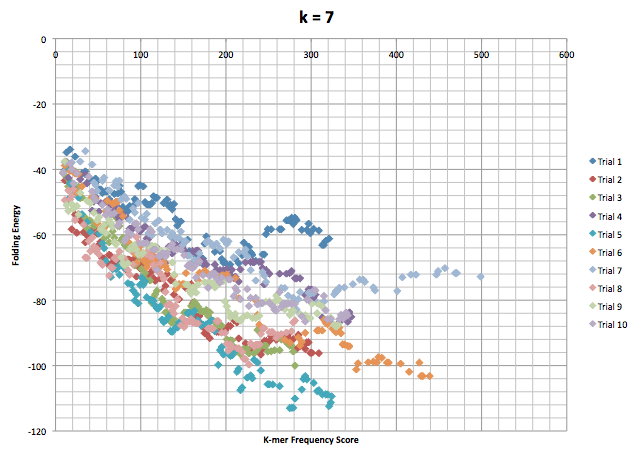
This next graph is for when the kmer length was set to 7. It is also 10 different runs of the program with 100 accepted mutations each.
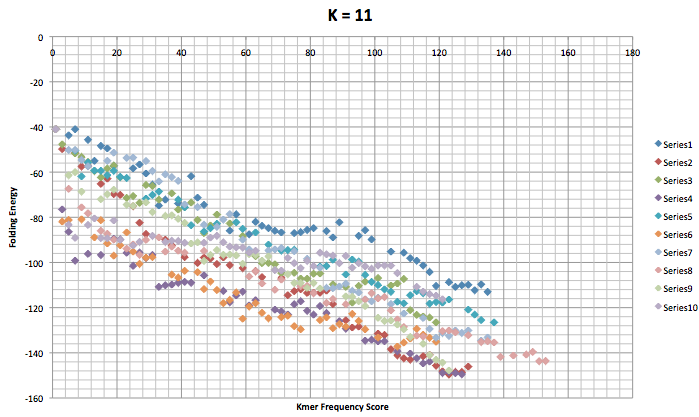
This next graph is for when the kmer length was set to 11. This graph is slightly different because it shows 10 trials with 50 mutations. It has less mutations because of the problems mention at the beginning of this post. One of my goals for this week is to get 100 mutations for each trial for k = 11 through k = 15 if possible,
In addition to running more trials with the current program I have, I will also be writing a similar program that in a way does the opposite of what the current one does. Instead of maximizing or minimizing the kmer frequency score I will write it so that it minimizes or maximizes the folding energy. It will then also calculate the kmer frequency score of the accepted mutations. Additionally, I will be working with JR to continue the experiments he has been running. We are currently looking into D-Tailor.